|
Petr Pancoska, PhD. |
|
Center for clinical pharmacology
|
|
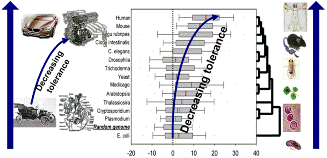
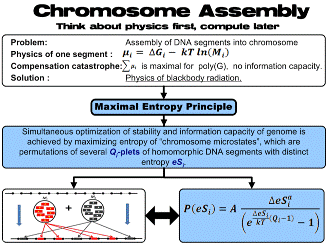
Entromic enthalpy, eH, represents the stability of the isolated DNA segment in question and entromic entropy represents a new and unique entromics discovery: eS = – kTln(M), M being the number of daughter DNA’s, determined uniquely from the sequenced genome parts. eS quantifies how flexible or “tolerant” is the genome neighborhood to the segment presence and its possible changes. We have extensive results, which provide direct validation of the entromics principles when applied to actual biological problems.
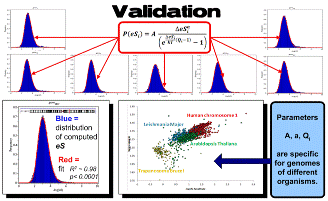
Examples: The basic law of entromics provides the first explicit and quantitative “ab initio” formulation of the mechanism behind the role of silent mutations in the genome properties optimization. The entropy eS quantifies the extent of compensation of incorporation energy of all genome parts during the cooperative evolution of multiple inter-related functions in biological system. This implies that coding sequences of the human genome are expected to be extensively eS-optimized. This prediction was validated by studying the distribution of differences between the eS computed for genomes of diverse organisms.
3. Applied Entromics. We have also derived the quantitative thermodynamic characterization for the assembly of many of these individual DNA segments into the chromosome. This process results in entromics' most important discovery, that a chromosome has to contain several (more than one) “daughter” segments with several diverse energies. |


|
About Entromics Patent information |
